Barkan Lab


- Alice Barkan
- Professor of Biology
- B.S., Massachusetts Institute of Technology
- Ph.D., University of Wisconsin
- Member of: Institute of Molecular Biology
- Office: 255A Klamath Hall
- Telephone: 541-346-5145
- Lab: 255 Klamath Hall
- Telephone: 541-346-2546
Research Interests
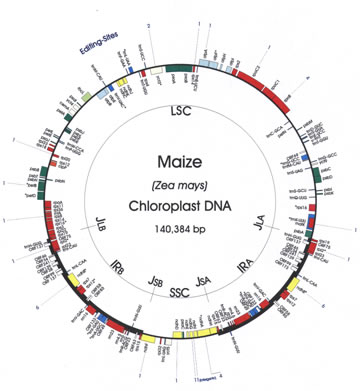 Research in the Barkan lab is directed at understanding how the genetic machineries in the chloroplast and nucleus communicate to produce a chloroplast that is responsive to environmental and developmental cues. We study mechanisms by which nuclear genes influence the synthesis of chloroplast proteins, with a focus on the post-transcriptional control of gene expression. We also use the chloroplast as a system in which to explore fundamental aspects of RNA/protein interactions including protein-facilitated group II intron splicing, and the mechanisms employed by the pentatricopeptide repeat (PPR) family, a remarkable family of RNA binding proteins that plays a multitude of roles in mitochondrial and chloroplast gene expression. In a recent collaborative project, we are exploring the translational dynamics underlying leaf and chloroplast development. Our research combines genetic and biochemical approaches to identify relevant genes, define their in vivo functions, and elucidate the underlying mechanisms. Tools we have developed for this work include: (i) the Photosynthetic Mutant Library (PML), a saturated collection of Mu transposon-induced non-photosynthetic maize mutants; (ii) Mu-Illumina, a method that allows us to rapidly identify the Mu transposon insertion underlying phenotypes-of-interest in high copy Mu-active maize lines; (iii) a RIP-chip assay that allows us to identify the native RNA ligands of chloroplast RNA binding proteins; and (iv) POGsDB, a database that groups Arabidopsis and rice proteins into Putative Orthologous Groups (POGs) and displays key features of each POG in a user-friendly format.
Research in the Barkan lab is directed at understanding how the genetic machineries in the chloroplast and nucleus communicate to produce a chloroplast that is responsive to environmental and developmental cues. We study mechanisms by which nuclear genes influence the synthesis of chloroplast proteins, with a focus on the post-transcriptional control of gene expression. We also use the chloroplast as a system in which to explore fundamental aspects of RNA/protein interactions including protein-facilitated group II intron splicing, and the mechanisms employed by the pentatricopeptide repeat (PPR) family, a remarkable family of RNA binding proteins that plays a multitude of roles in mitochondrial and chloroplast gene expression. In a recent collaborative project, we are exploring the translational dynamics underlying leaf and chloroplast development. Our research combines genetic and biochemical approaches to identify relevant genes, define their in vivo functions, and elucidate the underlying mechanisms. Tools we have developed for this work include: (i) the Photosynthetic Mutant Library (PML), a saturated collection of Mu transposon-induced non-photosynthetic maize mutants; (ii) Mu-Illumina, a method that allows us to rapidly identify the Mu transposon insertion underlying phenotypes-of-interest in high copy Mu-active maize lines; (iii) a RIP-chip assay that allows us to identify the native RNA ligands of chloroplast RNA binding proteins; and (iv) POGsDB, a database that groups Arabidopsis and rice proteins into Putative Orthologous Groups (POGs) and displays key features of each POG in a user-friendly format.
Barkan Lab - 255 Klamath Hall - abarkan@uoregon.edu